Twenty-one people in eight states infected with outbreak strain of Salmonella Heidelberg.
November 30, 2016
The Centers for Disease Control & Prevention (CDC) is working with Wisconsin health, agriculture and laboratory agencies, several other states and the U.S. Department of Agriculture's Animal & Plant Health Inspection Service (APHIS) to investigate a multistate outbreak of multi-drug-resistant Salmonella Heidelberg infections.
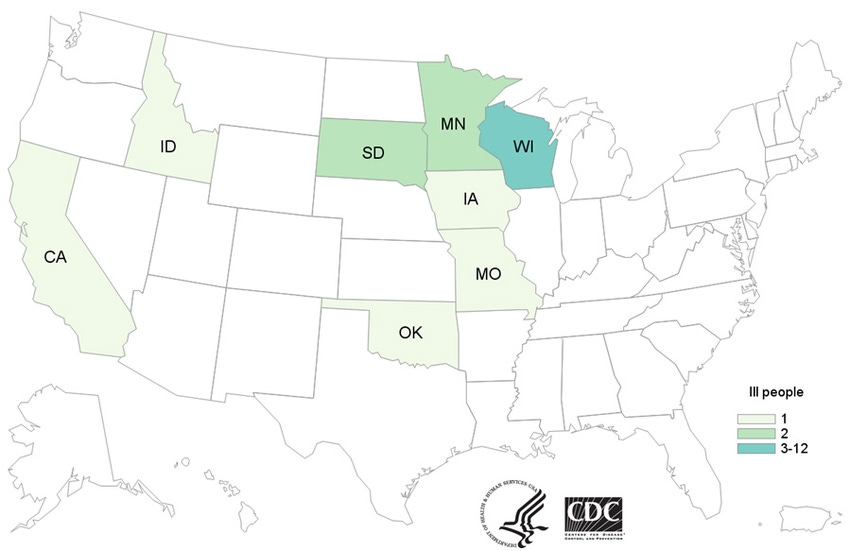
Twenty-one people infected with an outbreak strain of S. Heidelberg have been reported from eight states: Wisconsin, Iowa, Minnesota, Idaho, Missouri, Oklahoma, South Dakota and California. Among 19 people with available information, illnesses started on dates ranging from Jan. 11, 2016, to Oct. 24, 2016.
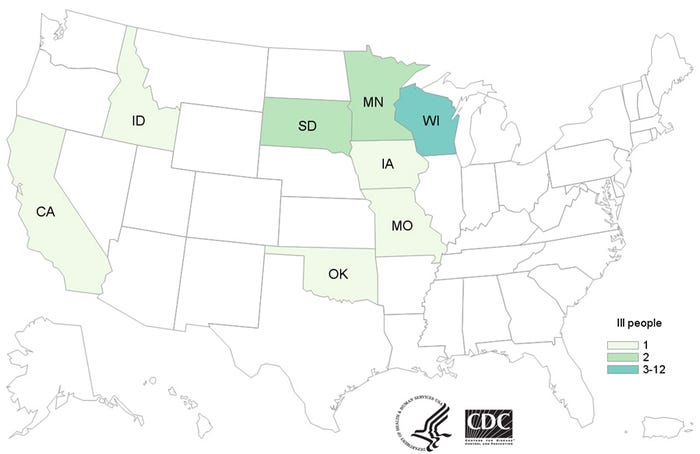
Source: CDC
Public health investigators used the PulseNet system to identify illnesses that may have been part of this outbreak. PulseNet, coordinated by CDC, is the national subtyping network of public health and food regulatory agency laboratories. PulseNet performs DNA fingerprinting on salmonella bacteria isolated from ill people by using techniques called pulsed-field gel electrophoresis (PFGE) and whole-genome sequencing (WGS). PulseNet manages a national database of these DNA fingerprints to identify possible outbreaks.
WGS showed that isolates from ill people are closely related to one another genetically. This close genetic relationship means that people affected by this outbreak are more likely to share a common source of infection.
Investigation of the outbreak
Epidemiologic, traceback and laboratory findings have identified dairy bull calves from livestock markets in Wisconsin as the likely source of infections. The claves have also been purchased for use with 4-H projects.
In interviews, ill people answered questions about any contact with animals and foods eaten in the week before becoming ill. Of the 19 people interviewed, 15 (79%) reported contact with dairy bull calves or other cattle. Some of the ill people interviewed reported that they became sick after their dairy bull calves became ill or died.
One ill person's dairy calves were tested for the presence of salmonella bacteria. This laboratory testing identified S. Heidelberg in the calves. Further testing using WGS showed that isolates from ill people are closely related genetically to isolates from these calves. This close genetic relationship means that the human infections in this outbreak are likely linked to ill calves.
As part of routine surveillance, the Wisconsin State Laboratory of Hygiene -- one of seven regional labs affiliated with CDC's Antibiotic Resistance Laboratory Network -- conducted antibiotic resistance testing on clinical isolates from the ill people associated with this outbreak. These isolates were found to be resistant to antibiotics and shared the same DNA fingerprints, showing that the isolates were likely related to one another.
WGS identified multiple antimicrobial resistance genes in outbreak-associated isolates from 15 ill people and eight cattle. This correlated with results from standard antibiotic resistance testing methods used by CDC's National Antimicrobial Resistance Monitoring System laboratory on clinical isolates from two ill people in this outbreak. The two isolates tested were susceptible to gentamicin, azithromycin and meropenem. Both were resistant to amoxicillin-clavulanic acid, ampicillin, cefoxitin, ceftriaxone, chloramphenicol, nalidixic acid, streptomycin, sulfisoxazole, tetracycline and trimethoprim-sulfamethoxazole and had reduced susceptibility to ciprofloxacin.
Antibiotic resistance limits treatment options and has been associated with increased risk of hospitalization, bloodstream infections and treatment failures in patients.
Traceback information available at this time indicates that most calves in this outbreak originated in Wisconsin, CDC said. Wisconsin health and agriculture officials continue to work with other states to identify herds that may be affected.
You May Also Like

.png?width=300&auto=webp&quality=80&disable=upscale)

