thumbnail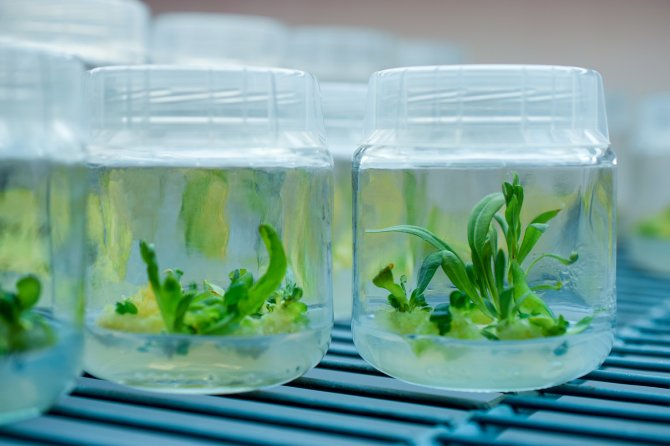
Agribusiness News
Researchers to examine how plants respond to touchResearchers to examine how plants respond to touch
Wageningen researchers will lead a large-scale research project into the sense of touch in plants over the next decade.
Subscribe to Our Newsletters
Feedstuffs is the news source for animal agriculture


.png?width=700&auto=webp&quality=80&disable=upscale)

.png?width=700&auto=webp&quality=80&disable=upscale)
.png?width=300&auto=webp&quality=80&disable=upscale)


.png?width=300&auto=webp&quality=80&disable=upscale)
.png?width=300&auto=webp&quality=80&disable=upscale)


.png?width=300&auto=webp&quality=80&disable=upscale)






.png?width=300&auto=webp&quality=80&disable=upscale)

.png?width=300&auto=webp&quality=80&disable=upscale)


.png?width=300&auto=webp&quality=80&disable=upscale)



.png?width=300&auto=webp&quality=80&disable=upscale)

.png?width=300&auto=webp&quality=80&disable=upscale)



